目的:
- 利用
seaborn畫出棒形圖(barplot),比較不同物種的編碼基因(coding genes)數量 - 在每根柱子上顯示計數值(count)
準備數據:
根據基因組數據庫(Ensembl release 104,NCBI)及基因組定序文獻,整理出以下脊椎動物的編碼基因數量:
| 物種 | 物種 (英文) | 學名 | 基因組 | 編碼基因數量 | 參考資料 |
|---|---|---|---|---|---|
| 小鼠 | mouse | Mus musculus | GRCm39 | 22,468 | Ensembl[1], Genome Reference Consortium (GRC)[2] |
| 人 | human | Homo sapiens | GRCh38.p13 | 20,442 | Ensembl[3], GRC[2] |
| 家犬 | dog | Canis lupus familiaris | CanFam3.1 | 20,257 | Ensembl[4], Wang et al. Commun Biol (2021)[5] |
| 家貓 | cat | Felis catus | Felis_catus_9.0 | 19,588 | Ensembl[6], Buckley et al. PLoS Genet (2020)[7] |
| 鴨嘴獸 | platypus | Ornithorhynchus anatinus | mOrnAna1.p.v1 | 17,418 | Ensembl[8], Zhou et al. Nature (2021)[9] |
| 雞 | chicken | Gallus gallus | GRCg6a | 16,878 | Ensembl[10], GRC[2] |
| 鴿 | pigeon | Columba livia | Cliv_2.1 | 15,392 | NCBI[11], Holt et al. G3 (2018)[12] |
| 鴯鶓 | emu | Dromaius novaehollandiae | droNov1 | 15,615 | Ensembl[13], Sackton et al. Science (2019)[14] |
| 鱷魚 | crocodile | Crocodylus porosus | Cpor_3.0 | 23,242 | NCBI[15], Ghosh et al. Genome Biol Evol (2020)[16] |
| 鱉 | softshell turtle | Pelodiscus sinensis | PelSin_1.0 | 18,189 | Ensembl[17], Wang et al. Nat Genet (2013)[18] |
| 喙頭蜥 | tuatara | Sphenodon punctatus | ASM311381v1 | 17,648 | Ensembl[19], Gemmell et al. Nature (2020)[20] |
| 蠑螈 | axolotl | Ambystoma mexicanum | AmbMex60DD | 23,251 | NCBI[21], Nowoshilow et al. Nature (2018)[22] |
| 熱帶爪蟾 | western clawed frog | Xenopus tropicalis | Xenopus_tropicalis_v9.1 | 19,987 | Ensembl[23], Xenbase[24] |
| 肺魚 | lungfish | Neoceratodus forsteri | neoFor_v3 | 31,120 | NCBI[25], Meyer et al. Nature (2021)[26] |
| 腔棘魚 | coelacanth | Latimeria chalumnae | LatCha1 | 19,569 | Ensembl[27], Amemiya et al. Nature (2013)[28] |
| 斑馬魚 | zebrafish | Danio rerio | GRCz11 | 25,592 | Ensembl[29], GRC[2] |
| 河豚 | fugu | Takifugu rubripes | fTakRub1.2 | 21,411 | Ensembl[30] |
| 海馬 | seahorse | Hippocampus comes | H_comes_QL1_v1 | 20,852 | Ensembl[31], Lin et al. Nature (2016)[32] |
| 鯊 | bamboo-shark | Chiloscyllium punctatum | Cpunctatum_v1.0 | 34,038 | NCBI[33], Hara et al. Nat Ecol Evol (2018)[34] |
| 盲鰻 | hagfish | Eptatretus burgeri | Eburgeri_3.2 | 16,513 | Ensembl[35], Yamaguchi et al. bioRxiv (2020)[36] |
| 七鰓鰻 | lamprey | Petromyzon marinus | Pmarinus_7.0 | 10,415 | Ensembl[37], Smith et al. Nat Genet (2013)[38] |
dict_CodingGeneNumber = {
'mouse': [22468, 'mammal'],
'human': [20442, 'mammal'],
'dog': [20257, 'mammal'],
'cat': [19588, 'mammal'],
'platypus': [17418, 'mammal'],
'chicken': [16878, 'bird'],
'pigeon': [15392, 'bird'],
'emu': [15615, 'bird'],
'crocodile': [23242, 'reptile'],
'softshell turtle': [18189, 'reptile'],
'tuatara': [17648, 'reptile'],
'axolotl': [23251, 'amphibian'],
'western clawed frog': [19987, 'amphibian'],
'lungfish': [31120, 'lobe-finned fish'],
'coelacanth': [19569, 'lobe-finned fish'],
'zebrafish': [25592, 'ray-finned fish'],
'fugu': [21411, 'ray-finned fish'],
'seahorse': [20852, 'ray-finned fish'],
'bambooshark': [34038, 'cartilaginous fish'],
'hagfish': [16513, 'jawless fish'],
'lamprey': [10415, 'jawless fish']
}
# 把dictionary轉換成dataframe
import pandas as pd
df = pd.DataFrame(data = dict_CodingGeneNumber,
index = ['count_CodingGenes', 'group']).T
# 把dataframe轉換成seaborn作圖格式
df.reset_index(inplace = True)
df.columns = ['species'] + list(df.columns[1:])
df.head(8)
DataFrame首八行:
species count_CodingGenes group
0 mouse 22468 mammal
1 human 20442 mammal
2 dog 20257 mammal
3 cat 19588 mammal
4 platypus 17418 mammal
5 chicken 16878 bird
6 pigeon 15392 bird
7 emu 15615 bird
代碼:
剛剛發現在matplotlib新版本3.4中,可直接在棒上方標註記數值(若使用舊版本,則需先取得每根柱子在圖中的位置,再從而計算出標記計數值文字的位置,比較麻煩)。先更新作圖模組:
python3 -m pip install matplotlib==3.4.3
python3 -m pip install seaborn==0.11.2
利用seaborn畫出基礎圖:
sns.barplot(data = df,
x = 'species', y = 'count_CodingGenes')

美化圖像:
- 改變圖像大小
- 柱子根據
'group'列資訊填色 - 圖例放在主圖框之外
- 更改橫、縱軸文字方向、大小
- 更改字型
# Set figure size
plt.figure(figsize = (18,7.5))
# Plot barplot
ax = sns.barplot(data = df,
x = 'species', y = 'count_CodingGenes',
hue = 'group', dodge = False,
palette = 'Set3')
# Legend box outside the main frame
plt.legend(bbox_to_anchor = (1.05, 1),
loc = 'upper left',
fontsize = 16)
# Rotate x-axis tick labels and change fontsize
plt.xticks(rotation=90, fontsize = 18)
plt.yticks(fontsize = 18)
# Rename axis labels
plt.xlabel('species', fontweight = 'bold', fontsize = 20)
plt.ylabel('number of coding genes', fontweight = 'bold', fontsize = 20)
# Set font
plt.rcParams["font.family"] = "Arial"
# Show plot
plt.tight_layout()
plt.show()

加上計數值[39]:
# Annotate counts
n_groups = len(df.group.unique())
for i in range(n_groups):
ax.bar_label(ax.containers[i], fontsize = 14)
# 或者以下方法
# Annotate counts
for container in ax.containers:
ax.bar_label(container, fontsize = 14)
註:如果只用ax.bar_label(ax.containers[0]),則只有第一組(最左方淺綠色柱子)被標記計數值。ax.containers中有八組數據。
完成圖:

參考資料及文獻
(All websites accessed on 2021-08-24)
- https://asia.ensembl.org/Mus_musculus/Info/Annotation.
- https://www.ncbi.nlm.nih.gov/grc.
- https://asia.ensembl.org/Homo_sapiens/Info/Annotation.
- https://asia.ensembl.org/Canis_lupus_familiaris/Info/Annotation.
- Wang, C. et al. A novel canine reference genome resolves genomic architecture and uncovers transcript complexity. Commun Biology 4, 185 (2021).
- https://asia.ensembl.org/Felis_catus/Info/Annotation.
- Buckley, R. M. et al. A new domestic cat genome assembly based on long sequence reads empowers feline genomic medicine and identifies a novel gene for dwarfism. PLoS Genet 16, e1008926 (2020).
- https://asia.ensembl.org/Ornithorhynchus_anatinus/Info/Annotation.
- Zhou, Y. et al. Platypus and echidna genomes reveal mammalian biology and evolution. Nature 592, 756–762 (2021).
- https://asia.ensembl.org/Gallus_gallus/Info/Annotation.
- https://www.ncbi.nlm.nih.gov/assembly/1489441.
- Holt, C. et al. Improved genome assembly and annotation for the rock pigeon (Columba livia). G3 Genes Genomes Genetics 8, g3.300443.2017 (2018).
- https://asia.ensembl.org/Dromaius_novaehollandiae/Info/Annotation.
- Sackton, T. B. et al. Convergent regulatory evolution and loss of flight in paleognathous birds. Science 364, 74–78 (2019).
- https://www.ncbi.nlm.nih.gov/assembly/GCA_000768395.2#/st.
- Ghosh, A. et al. A high-quality reference genome assembly of the saltwater crocodile, Crocodylus porosus, reveals patterns of selection in Crocodylidae. Genome Biol Evol 12, 3635-3646 (2020).
- https://asia.ensembl.org/Pelodiscus_sinensis/Info/Annotation.
- Wang, Z. et al. The draft genomes of soft-shell turtle and green sea turtle yield insights into the development and evolution of the turtle-specific body plan. Nat Genet 45, 701–706 (2013).
- https://asia.ensembl.org/Sphenodon_punctatus/Info/Annotation.
- Gemmell, N. J. et al. The tuatara genome reveals ancient features of amniote evolution. Nature 584, 403–409 (2020).
- https://www.ncbi.nlm.nih.gov/genome/381.
- Nowoshilow, S. et al. The axolotl genome and the evolution of key tissue formation regulators. Nature 554, 50–55 (2018).
- https://asia.ensembl.org/Xenopus_tropicalis/Info/Annotation.
- Karimi, K. et al. Xenbase: a genomic, epigenomic and transcriptomic model organism database. Nucleic Acids Res 46, D861–D868 (2018).
- https://www.ncbi.nlm.nih.gov/genome/7137?genome_assembly_id=1532268.
- Meyer, A. et al. Giant lungfish genome elucidates the conquest of land by vertebrates. Nature 590, 284–289 (2021).
- https://asia.ensembl.org/Latimeria_chalumnae/Info/Annotation.
- Amemiya, C. T. et al. The African coelacanth genome provides insights into tetrapod evolution. Nature 496, 311–316 (2013).
- https://asia.ensembl.org/Danio_rerio/Info/Annotation.
- https://asia.ensembl.org/Takifugu_rubripes/Info/Annotation.
- https://asia.ensembl.org/Hippocampus_comes/Info/Annotation.
- Lin, Q. et al. The seahorse genome and the evolution of its specialized morphology. Nature 540, 395–399 (2016).
- https://www.ncbi.nlm.nih.gov/genome/12366?genome_assembly_id=397945.
- Hara, Y. et al. Shark genomes provide insights into elasmobranch evolution and the origin of vertebrates. Nat Ecol Evol 2, 1761–1771 (2018).
- https://asia.ensembl.org/Eptatretus_burgeri/Info/Annotation
- Yamaguchi, K. et al. Inference of a genome-wide protein-coding gene set of the inshore hagfish Eptatretus burgeri. bioRxiv (2020). doi:10.1101/2020.07.24.218818
- https://asia.ensembl.org/Petromyzon_marinus/Info/Annotation
- Smith, J. J. et al. Sequencing of the sea lamprey (Petromyzon marinus) genome provides insights into vertebrate evolution. Nat Genet 45, 415–421 (2013).
- https://stackoverflow.com/questions/55104819/display-count-on-top-of-seaborn-barplot
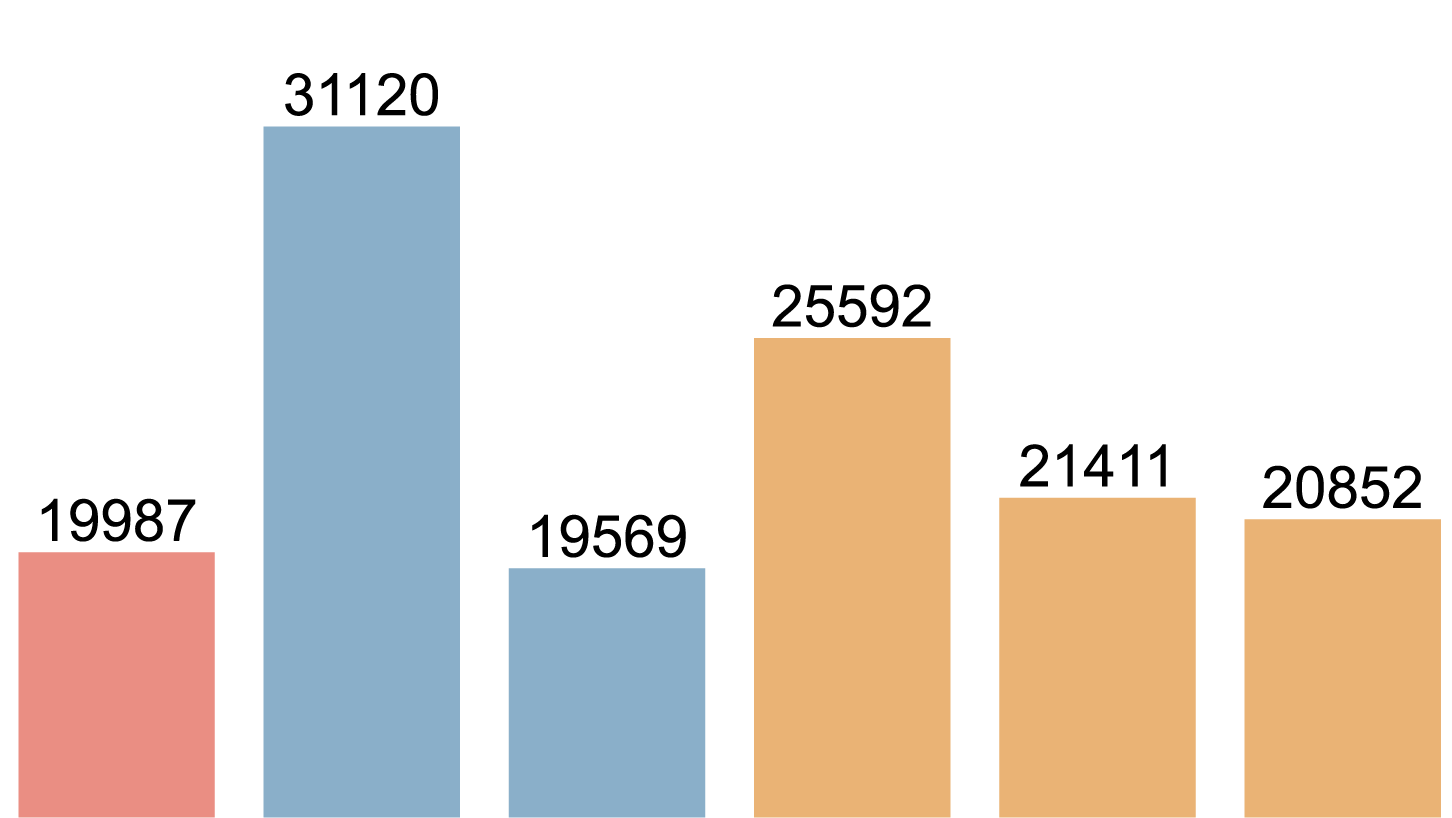
Leave a Reply